Plot community-based enrichment results for a filtered GRN object
plotCommunitiesEnrichment.RdSimilarly to plotGeneralEnrichment and plotTFEnrichment, the results of the community-based enrichment analysis are plotted.
This function produces multiple plots. First, one plot per community to summarize the community-specific enrichment.
Second, a summary heatmap of all significantly enriched terms across all communities and for the whole eGRN. The latter allows to compare the results with the general network enrichment.
Third, a subset of the aforementioned heatmap, showing only the top most significantly enriched terms per community and for the whole eGRN (as specified by nID) for improved visibility
plotCommunitiesEnrichment(
GRN,
outputFolder = NULL,
basenameOutput = NULL,
selection = "byRank",
communities = NULL,
topn_pvalue = 30,
p = 0.05,
nSignificant = 2,
nID = 10,
maxWidth_nchar_plot = 50,
display_pAdj = FALSE,
plotAsPDF = TRUE,
pdf_width = 12,
pdf_height = 12,
pages = NULL,
forceRerun = FALSE
)Arguments
- GRN
Object of class
GRN- outputFolder
Character or
NULL. DefaultNULL. If set toNULL, the default output folder as specified when initiating the object ininitializeGRNwill be used. Otherwise, all output from this function will be put into the specified folder. If a folder is provided, while we recommend specifying an absolute path, a relative one also works.- basenameOutput
NULLor character. DefaultNULL. Basename of the output files that are produced. If set toNULL, a default basename is chosen. If a custom basename is specified, all output files will have the chosen basename as file prefix, be careful with not overwriting already existing files (ifforceRerunis set toTRUE)- selection
Character. Default
"byRank". One of:"byRank","byLabel". Specify whether the communities will be selected based on their rank or explicitly by their label. Note that the label is independent of the rank. When set to"byRank", the largest community (with most vertices) always has a rank of 1.- communities
NULLor numeric vector or character vector. DefaultNULL. If set toNULL, all community enrichments that have been calculated before are plotted. If a numeric vector is specified (whenselection = "byRank"), the rank of the communities is specified. For example,communities = c(1,4)then denotes the first and fourth largest community. If a character vector is specified (whenselection = "byLabel"), the name of the communities is specified instead. For example,communities = c("1","4")then denotes the communities with the names "1" and "4", which may or may not be the largest and fourth largest communities among all.- topn_pvalue
Numeric. Default 30. Maximum number of ontology terms that meet the p-value significance threshold to display in the enrichment dot plot
- p
Numeric. Default 0.05. p-value threshold to determine significance.
- nSignificant
Numeric > 0. Default 3. Threshold to filter out an ontology term with less than
nSignificantoverlapping genes.- nID
Numeric > 0. Default 10. For the reduced summary heatmap, number of top terms to select per community / for the general enrichment.
- maxWidth_nchar_plot
Integer (>=10). Default 50. Maximum number of characters for a term before it is truncated.
- display_pAdj
TRUEorFALSE. DefaultFALSE. Is the p-value being displayed in the plots the adjusted p-value? This parameter is relevant for KEGG, Disease Ontology, and Reactome enrichments, and does not affect GO enrichments.- plotAsPDF
TRUEorFALSE. DefaultTRUE.Should the plots be printed to a PDF file? If set toTRUE, a PDF file is generated, the name of which depends on the value ofbasenameOutput. If set toFALSE, all plots are printed to the currently active device. Note that most functions print more than one plot, which means you may only see the last plot depending on your active graphics device.- pdf_width
Number>0. Default 12. Width of the PDF, in cm.
- pdf_height
Number >0. Default 12. Height of the PDF, in cm.
- pages
Integer vector or
NULL. DefaultNULL. Page number(s) to plot. Can be used to plot only specific pages to a PDF or the currently active graphics device.- forceRerun
TRUEorFALSE. DefaultFALSE. Force execution, even if the GRN object already contains the result. Overwrites the old results.
Value
The same GRN object, without modifications.
Examples
# See the Workflow vignette on the GRaNIE website for examples
GRN = loadExampleObject()
#> Downloading GRaNIE example object from https://git.embl.de/grp-zaugg/GRaNIE/-/raw/master/data/GRN.rds
#> INFO [2023-08-16 17:29:10] Storing GRN@data$RNA$counts matrix as sparse matrix because fraction of 0s is > 0.1 (0.44)
#> Finished successfully. You may explore the example object. Start by typing the object name to the console to see a summaty. Happy GRaNIE'ing!
GRN = plotCommunitiesEnrichment(GRN, plotAsPDF = FALSE, pages = 1)
#> INFO [2023-08-16 17:29:10] Including terms only if overlap is at least 2 genes.
#> INFO [2023-08-16 17:29:10] Plotting the enrichment for the following communities: 1,5,4,2,3,6
#> INFO [2023-08-16 17:29:11] Plotting directly
#> INFO [2023-08-16 17:29:11] Plotting results for ontology GO_BP
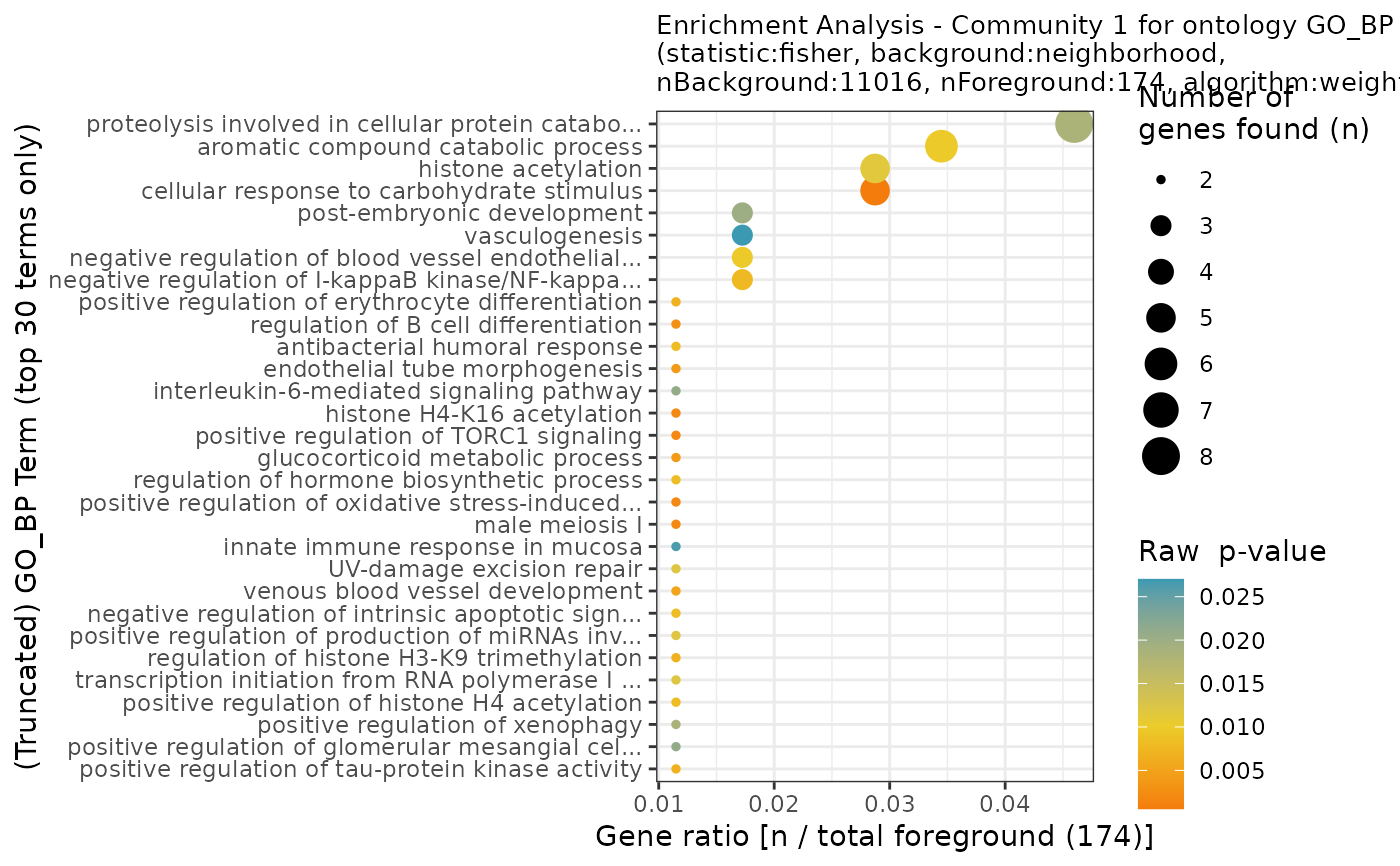 #> INFO [2023-08-16 17:29:12] Plotting results for ontology GO_MF
#> INFO [2023-08-16 17:29:12] Finished successfully. Execution time: 1.8 secs
#> INFO [2023-08-16 17:29:12] Plotting results for ontology GO_MF
#> INFO [2023-08-16 17:29:12] Finished successfully. Execution time: 1.8 secs